Full Text
The global burden of tuberculosis (TB) has been declining slowly over the past several years, but it remains high [1]. It is still a global epidemic, especially in developing countries. Fifty-eight percent of the total TB cases occur in South-East Asia and Western Pacific regions. India stands first with highest number of cases closely followed by China, South Africa, Indonesia and Pakistan. India accounts for one fifth of the global incidence [2]. The problem is overwhelmed with “The Deadly Trio” comprising human immunodeficiency virus (HIV), tuberculosis and diabetes mellitus. One person dies of Tuberculosis every two minutes in India according to World Health Organization (WHO) report in 2014.
TB is a multisystem disease; it can affect any organ or tissue excluding hair and nails. Even today pulmonary tuberculosis accounts for majority of cases and is the main transmissible form of disease. However, TB affecting other areas of body also contributes to burden of disease and does not receive specific attention in international control strategies [3].
A rapid and accurate diagnosis is helpful to ensure cure of the infected patient, prevent emergence of resistant strains, interrupt person-to-person transmission and implement effective public health interventions. The need for an efficient Tuberculosis diagnostic becomes evident from the fact that for every patient of Tuberculosis who can be detected using microscopy, nine have to be screened due to low sensitivity of microscopy [4].
The most widely used method to detect pulmonary TB is the 125-year-old sputum smear microscopy test, which has a number of drawbacks, including low sensitivity (especially in HIV-positive individuals and children) [5]. Deficiency of sample received and hence a yield of very few bacilli is the difficulty with non-sputum samples. This further reduces the sensitivity of acid- fast bacilli (AFB) smear and culture of such samples. Chest radiography used for smear negative cases is less specific for tuberculosis. Culture remains the gold standard but it is time consuming and not always available.
Molecular methods have the potential to detect both M. tuberculosis and non-tuberculous mycobacteria directly from clinical samples [6], one among them being polymerase chain reaction (PCR). The potential of PCR as a diagnostic test for tuberculosis has been investigated in a large number of studies [7-10]. In many studies, problems with false-positive PCR results, at rates ranging from 0.8% to 30% have been reported. Specificity of PCR results varies between laboratories due to procedural differences, differences in cross-contamination rates and the choice of primers [11].
This study was designed to compare ZN staining, culture and PCR methods in detecting Mycobacterium tuberculosis in non-sputum samples.
Material and methods
Study area: The study was conducted in the Department of Microbiology & Molecular Biology, Krishna Institute of Medical Sciences (KIMS), Secunderabad, a tertiary care centre.
Study population: All pulmonary non-sputum samples received for M. tuberculosis culture from clinically suspected cases of all ages were included in the study.
Study duration: One year (1 July 2015 to 31 June 2016).
Study design: Sample processing and analysis of data in Microbiology and Molecular Biology departments was done.
A total of 199 non-sputum samples were received during the one year time period which included 104 bronchial wash samples, 35 pleural fluids, 21 pus samples, 18 bronchoalveolar lavage (BAL) fluid, 12 ultra sound or CT guided fluid and 9 tissue samples. Bronchial wash, BAL and tissue samples were sent without any preservative added.
Inclusion criteria: (a) All pulmonary non-sputum samples, (b) Both inpatients and out patients, (c) All age groups of patients
Exclusion criteria: (a) Sputum samples, (b) Samples received from other organ systems (CSF, Urine, Blood and Feces), (c) Samples received for AFB smear alone
All the samples received were examined by routine smear microscopy and culture. PCR could be done for 133 patient samples only, other patients being non-affordable.
Sample processing, culture and PCR
Acid-fast staining: A direct smear was made from all the samples and stained by modified Ziehl-Neelsen technique and examined under oil-immersion (100X) using light microscope for the presence of acid fast bacilli. The typical cell morphology of M. tuberculosis as seen in carbolfuchsin – stained smears is a thin, slightly curved bacillus that measures 0.3 - 0.6 × 1 - 4 mm, deeply red staining (strongly acid-fast), with a distinct beaded appearance [12]. They may also appear as irregular aggregates or parallel strands. Smears were designated positive or negative for acid fast bacilli.
Smears for quality control prepared from TB culture positive samples were stained and reviewed at the start of work each day. Proficiency testing is done once in 15 days and Inter Lab Comparison once in three months.
Culture of specimen
The samples were inoculated on Lowenstein-Jensen (LJ) medium and incubated at 37°c. Inoculated slants were examined twice weekly for the first two weeks (for rapid growing mycobacteria, fungi and contaminating bacteria) and then once weekly thereafter for 6 weeks. Cultures that showed no growth after 6 weeks were considered negative. Growth observed was smeared and tested by ZN staining. Slow growing, non-pigmented, dry, rough, raised, irregular colony with wrinkled surface, which is niacin positive, showing acid fast bacilli on smear, was considered as M. tuberculosis. Niacin negative isolates were further identified based on their morphological and biochemical characters. Quality control was maintained as per recommendations.
PCR of clinical specimen: PCR was performed using 3B BIOTUB Kit (Biotools B&M labs, Spain) as a Nested PCR.The steps involved include DNA extraction, Amplification and Gel electrophoresis. The DNA extraction of samples was done as per manufacturer instructions. The sensitivity and specificity of the test is deemed high as it includes two amplification steps.
The first amplification reaction uses a pair of primers hybridizing with IS6110 region, a conserved sequence specific to M. tuberculosis complex. The second reaction is a nested amplification reaction using a second pair of primers, thus increasing the sensitivity of detection several-fold. The primer pair used in second step has a high Tm, which improves specificity of the reaction. The gel is then read under UV light after ethidium bromide staining. The system has an internal control plasmid DNA for identification of processed specimens containing any substances that could inhibit PCR.
Samples containing M. tuberculosis DNA render band of 219 base pairs (2nd PCR product) and internal control DNA renders a band of 571 base pairs. In rare samples with high load of bacilli, band of 371 base pairs (1st PCR product) could also be generated. Samples without internal control band were repeated and retested. Positive and negative controls were run with each cycle.
Statistical analysis
Analysis of the obtained data was done by calculating sensitivity, specificity, positive predictive value, negative predictive value and efficiency considering culture as the gold standard.
Results & discussion
In any hospital setting, majority of samples submitted for detection of tuberculosis are sputum samples. In order to emphasize the role of other samples, sputum samples were excluded from the study. The lower yield of bacilli due to irregular, intermittent release of bacteria in diseased individuals results in highly variable recovery pattern from respiratory specimen. The category wise distribution of samples is depicted in figures 1 & 2.
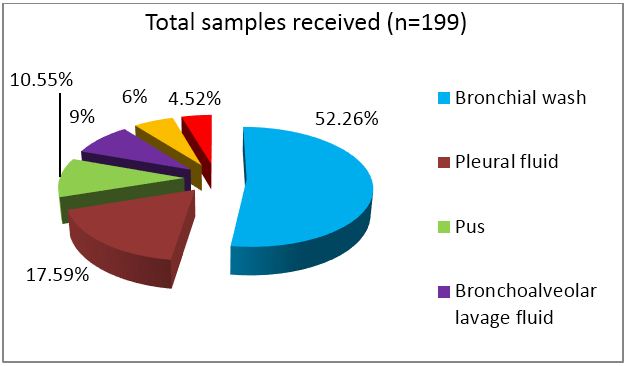
Figure 1: Category wise distribution of samples received.
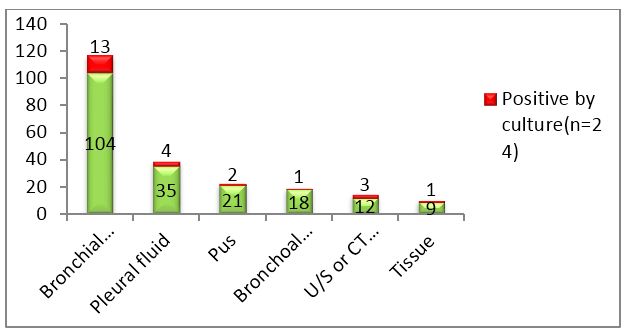
Figure 2: Column chart showing total samples received and number of culture positives in each category.
Out of 199 samples, 33(16.58%) were smear positive, 24(12.06%) were culture positive and 21(10.55%) were PCR positive. Out of 24 culture positives, 18 were sent for PCR. Only those samples were further analyzed.
The age distribution of tuberculosis is high in economically productive group as per WHO and the disease has male preponderance which could not be appreciated in this study. The age group distribution was quite remarkable as seen in figure 3. Although both genders presented in middle age (31-60 years), an early peak was observed in case of females.
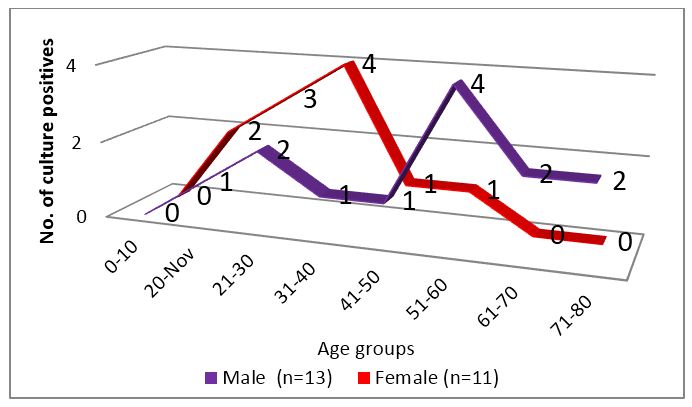
Figure 3: Age group distribution of culture positive patients.
Smears stained with ZN stain must be scanned with an oil-immersion (100X) objective. This limits the total area of slide that can be viewed in a given unit of time. There is a good correlation between positive AFB smear and positive culture if the disease is severe. In minimal or less-advanced cases the correlation is less. Images of culture positive samples are shown in figure 4.
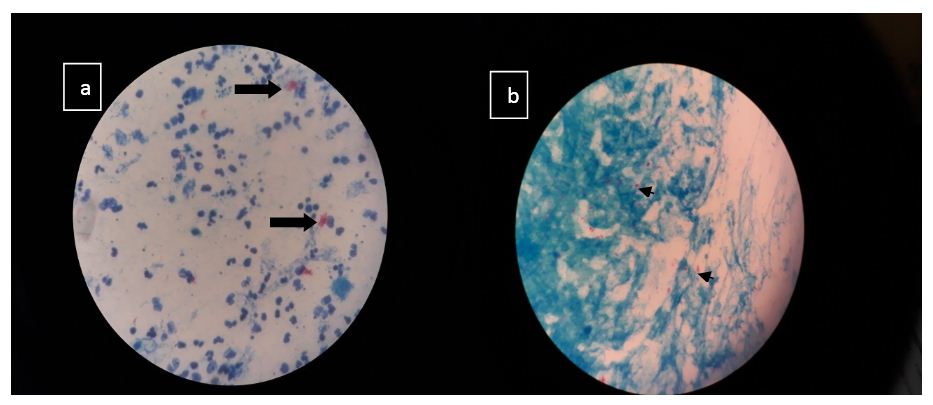
Figure 4: Image (a) Plenty of pus cells with plenty acid fast bacilli in clumps suggestive of M. tuberculosis in a pus sample Image (b) Occasional pus cell, scanty epithelial cells with plenty acid fast bacilli in singles and clumps suggestive of M. tuberculosis in a bronchial wash sample.
Lipsky, in a review of factors affecting the clinical value of microscopy for AFB, concluded that when the results of all specimens from each patient are considered in total, the acid-fast smear has a high predictive value and remains one of the rapidly performed tests [12].
False positives were more with smear microscopy as compared to PCR as shown in tables 1 & 2.
Table 1: Comparison of Ziehl-Neelsen smear microscopy with culture as the gold standard.
|
ZN smear/Culture
|
Culture positive
|
Culture negative
|
Total
|
|
Smear positive
|
11 (TP)
|
22 (FP)
|
33
|
|
Smear negative
|
13 (FN)
|
153 (TN)
|
166
|
|
Total
|
24
|
175
|
199
|
Abbreviations: TP - True positives, TN - True negatives, FN - False negatives, FP – False positives.
Table 2: Comparison of PCR with culture as the gold standard.
|
PCR/Culture
|
Culture positive
|
Culture negative
|
Total
|
|
PCR positive
|
12 (TP)
|
9 (FP)
|
21
|
|
PCR negative
|
6 (FN)
|
106 (TN)
|
112
|
|
Total
|
18
|
115
|
133
|
Abbreviations: TP - True positives, TN - True negatives, FN - False negatives, FP - False positives.
The usage of primer pair targeting IS6110 region, a conserved sequence specific to M. tuberculosis complex avoids genetic variability associated with antibiotic resistance [13]. False negatives and false positives detected with PCR are low when compared to smear microscopy. The PCR kit used does not differentiate between dead and live bacilli and among the different species of Mycobacteria, bringing down its specificity. The results are comparable to other similar studies [14]. Figure 5 shows positive and negative controls along with patient sample results in PCR.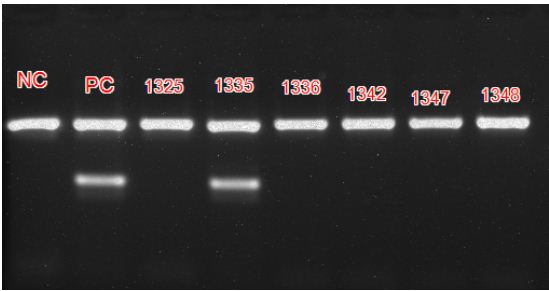
Figure 5: PCR result after ethidium bromide staining captured using UV light showing Positive and Negative controls and patient samples.
Foot note: Internal control band rendered at 571 base pairs and positive band at 219 base pairs. A negative control (NC), positive control (PC) and 6 patient samples with identification numbers are shown. Sample numbered 1335 is positive.
Comparative evaluation of different diagnostics is shown in table 3. Lower sensitivity of PCR with non-sputum samples, especially pleural fluid has already been demonstrated [15]. Though BAL fluid, bronchial wash, gastric lavage and aspiration perform reliably with PCR, pleural fluid is suboptimal. Pleural biopsy is the preferred sample.
Table 3: Comparative sensitivity, specificity, predictive value and efficiency of the two diagnostics with culture as the gold standard.
|
Diagnostic
|
Sensitivity (%)
|
Specificity (%)
|
Positive predictive value
|
Negative predictive value
|
Efficiency (%)
|
|
Microscopy
|
45.83 (11/24)
|
87.42 (22/175)
|
33.33 (11/33)
|
92.16 (153/166)
|
82.41 (164/199)
|
|
PCR
|
66.67 (12/18)
|
92.17 (106/115)
|
57.14 (12/21)
|
94.64 (106/112)
|
88.72 (118/133)
|
Conclusion
Though PCR attracted enormous attention, false positive results may mislead clinicians. Another limitation of PCR is its cost which is of prime concern in developing countries like India, where affordability is a factor. WHO endorsed techniques like cartridge-based molecular tests (Xpert MTB/RIF); line probe assays (HAIN’s MTBDR) which can simultaneously detect resistance pattern to certain anti-tubercular drugs directly from the clinical samples could further improve patient treatment. Molecular methods provide useful information quickly, but they cannot replace culture methods, at least in the near future. PCR and culture used together, in resourceful settings could fetch faster and more reliable results than any diagnostic test used alone, specifically in non-sputum samples.
Acknowledgements
I sincerely thank the whole department of Microbiology, Molecular Biology and Laboratory sciences, Krishna Institute of Medical sciences without whose support this work could not be carried out.
Conflicts of interest
Authors declare no conflicts of interest.
References
[1] World Health Organization. Global Tuberculosis Report. 2013.
[2] World Health Organization. Global tuberculosis control. 2010.
[3] Kulchavenya E. Extrapulmonary tuberculosis: Are statistical reports accurate? Ther Adv Infect Dis. 2014; 2(2):61-70.
[4] Modi Parekh K, Inamdar V, Jog A, Kar A. A comparative study of the diagnosis of pulmonary tuberculosis using conventional tools and polymerase chain reaction. Ind J Tuberc. 2006; 53(2):69-76.
[5] Centres for Disease Control and Prevention (CDC). Updated guidelines for the use of nucleic acid amplification tests in the diagnosis of tuberculosis. Morb Mortal Wkly Rep. 2009; 58:7–10.
[6] Soini H, Musser JM. Molecular diagnosis of mycobacteria. Clin Chem. 2001; 47(5):809-814.
[7] Watterson SA, and Drobniewski FA. Modern laboratory diagnosis of mycobacterial infections. J ClinPathol. 2000; 53(10):727-932.
[8] Brisson-Noel A, Gicquel B, Lecossier D, Lévy-Frébault V, Nassif X, et al. Rapid diagnosis of tuberculosis by amplification of mycobacterial DNA in clinical samples. Lancet. 1989; 4(8671):1069-1071.
[9] Rattan A. PCR for diagnosis of tuberculosis: Where are we now? Indian J Tuberc. 2000; 47:79-82.
[10] Foulds J, O Brien R. New tools for the diagnosis of tuberculosis: The perspective of developing countries. Int J Tuberc Lung Dis. 1998; 2(10): 778-783.
[11] Centers for Disease Control and Prevention (CDC). Update: Nucleic acid amplification tests for tuberculosis. Morb Mortal Wkly Rep. 2000; 49(26):593–594.
[12] Koneman EW, Janda WM, Hall GS, Church DL, Procop GW, et al. Mycobacteria Koneman’s color atlas and textbook of Diagnostic Microbiology. 2016; 7:1220-1260.
[13] Aryan E, Makvandi M, Farajzadeh A, Huygen K, Bifani P, et al. A novel and more sensitive loop-mediated isothermal amplification assay targeting IS6110 for detection of Mycobacterium tuberculosis complex. Microbiol Res. 2010; 165(3):211–220.
[14] Chakravorthy S, Sen MK, Tyagi JS. Diagnosis of extrapulmonary tuberculosis by smear, culture, and PCR using universal sample processing technology. J Clin Microbiol. 2005; 43(9):4357-4362.
[15] World Health organization. Policy update: Automated realtime nucleic acid amplification technology for rapid and simultaneous detection of tuberculosis and rifampicin resistance: Xpert MTB/ RIF system for the diagnosis of pulmonary and extrapulmonary TB in adults and children 2013.